Python and Databases
With all the cool things I recently discovered with Python, and some headaches with one of our systems at work, I wanted to make a case for setting up a dedicated Python Notebook server for my department. Before I get into the fun details of testing the setup for our needs, I should probably explain our needs. My goal is to replace our (very expensive) SPSS licenses with something not so expensive. It just so happens that Python, Pandas, MatplotLib, iPython and all the other goodies come at the correct price of Free. The only cost that should come out of this is the virtual server that would be needed to run it.
In order to evaluate if python could be used provide a viable alternative, I had to ask: what do we use SPSS for currently? Well, we load data from csv format to MS SQL and run an occasional statistical analysis. That's about it.
Preparing for Testing
The environment I needed to test for is a Microsoft one so I booted up my Windows 2012 Server in VirtualBox. Once again, I opted to use some of the NOAA data I had recently downloaded since it provided roughly the amount of data we will need to be able to load at once. I already had SQL Server 2014 installed so I just installed Python 2.8 via Anaconda, then added SQL Alchemy, pyodbc, and possibly one or two other dependencies that I missed writing down.
The NOAA file I used for testing contains a month's worth of precipitation data. The reason I chose that file was because our largest single-file loads are roughly 40 MB when in .csv format and are roughly 100 columns wide and 45,000 records long. The NOAA precipitation file is almost 33 MB so is close to total size on disk, but has a much different shape. It's almost 1.5 million records long and only 6 columns wide, but should work well enough for testing purposes.
Crash and Burn
It is safe to say that my first attempt to load the file to SQL didn't work so well. I ran the to_sql() method provided by Pandas on the full data set... and the VM promptly seized up, froze, and died. I should have known better because the default functionality is to take the whole file, load it to RAM, then try to load the entire chunk to SQL in one single batch. For this first attempt the virtual machine had 2GB RAM allocated which is almost nothing to Windows. So, I tried giving it 3GB and ran it again, which also failed miserably.
The solution was was simple, and only required setting the chunksize property when reading the csv file. I then was able to loop through the chunks and append each set of records to the database in batches.
Slow and Steady
After crashing my virtual machine twice, I wanted to make sure I was able to
get through the file at least once so I set the chunksize property to a very
cautious value of 1,000 and sat back with some popcorn. After telling it to
execute I closely watched the task manager in Windows. The python process was
consuming 80-90 MB RAM and taking up 5-15% of the CPU. It looked good, so I
went to SSMS to run a
COUNT(*)
on the table I was loading to, and was less excited. It was loading
less than 1,200 records per second which was lower than I expected to see. It
took almost 21 minutes to load all 1,488,000 records. Ok.... I'm still impressed
by being able to load almost 1.5 million records into MS SQL Server running
inside a Windows server on my 2+ year old Macbook Pro. That's cool. But... I'm
impatient! Also, I noticed that towards the beginning of processing, the SQL
Server process was taking under 200MB of RAM, but towards the end was up
around 800 MB of RAM. It was also hammering the CPU pretty hard, keeping it
between 60-90%. After the data was loaded that RAM wasn't released back to the
system either, SQL decided it was happier with it.
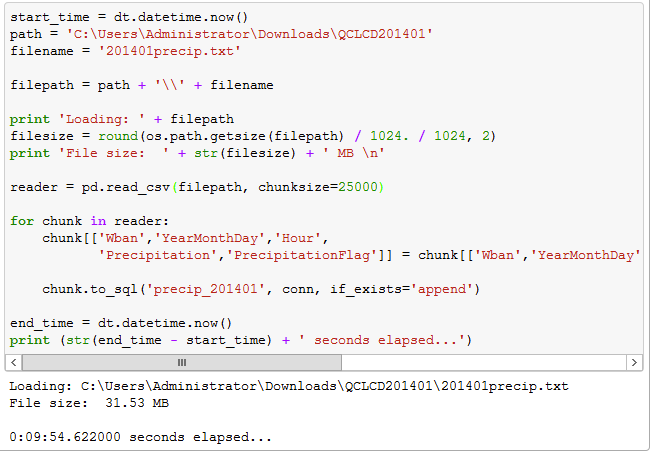
By setting the chunksize property up to 25,000 it reduced the load time to 9 minutes and 55 seconds. That was less than half the time than my first run, and loading roughly 2,500 records per second. The screenshot below shows the code I executed and the results showing file size and processing time.
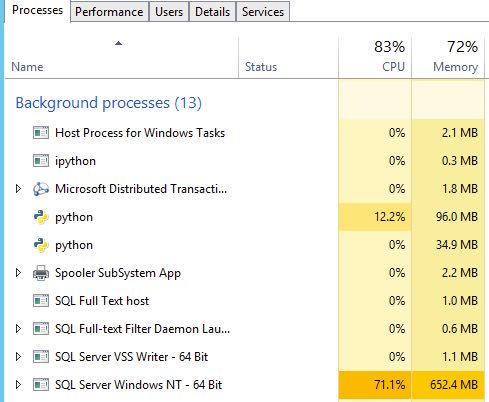
With the chunk size at 25k, the python process was now taking between 120 and 140 MB RAM and holding onto 20-30% of the CPU, but SQL Server was taking almost 1 GB of the 3GB given to the entire server, and was keeping at least 40% of the processor occupied, but hitting 100% more than a few times. At this point, I've decided that having SQL Server running on my little vm instead of a mini-monster with 128 GB RAM is the real bottleneck.
Performance Comparison
I won't lie, I was impressed by loading that many records to a high-demand server (MS SQL) in that little time on this little hardware... but I was curious how running the same import on a Debian server with PostgreSQL would stack up, given the same exact hardware. So I booted up my development Python virtual machine running Debian with one core and one GB RAM. I installed PostgreSQL 9.1 and prepared to run the same test to load a bunch of data.
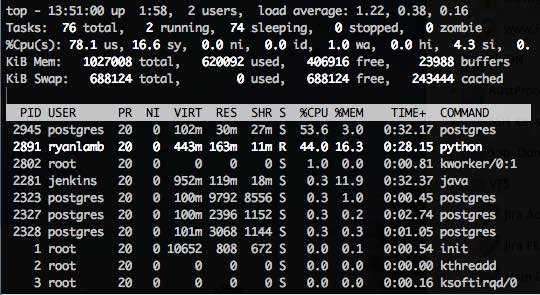
Watching the resources, I noticed that the python process on the Debian vm was taking about 165 MB RAM and keeping the CPU at about 30-50%. This is noticeably higher than on the Windows box. Then I looked at the postgres process which is taking 45-55% of the CPU... but only 40 MB RAM! Woah! The virtual machine only has 1 GB total RAM and isn't even using all of it to handle this load. The total time to load the same 1.5 million records came in at just over 6 minutes, with a load rate of 3,767 records per second. Well, that shows that my slow 5,800 rpm HDD wasn't the sole cause for the time, because PSQL with less RAM chopped off another 3.5 minutes.
Findings
For the purposes of what we need, I don't have concerns about how Python and Pandas will hold up in our production environment. As I mentioned in my prior post, I have already come to love NumPy, SciPy and MatPlotLib, so that covers the statistical analysis we have previously used SPSS for. It looks like it could work out well for us!
If you need to run Windows and/or MS SQL Server, you can get the job done. If you aren't tied to the MS stack though, I would recommend setting up a linux virtual machine, installing PostgreSQL and trying it out that way. My experience shows you'll get more power for the price with the lower overhead.
Have you had different experiences with using Python this way? Let me know in the comments!
